How do cells integrate actin-based motility and transcriptional regulation ?
Complex regulatory networks determine mammalian cell motility, morphology, and proliferation. Changes in actin dynamics through upstream signals and cell contacts activate a transcriptional program, which in turn itself affects cell behaviour through the induction of cytoskeletal and other components. We utilise a variety of methods from biochemistry, functional genomics and cell biology to understand these networks under normal physiological conditions, as well as their deregulation during pathogenesis, e.g. cancer.
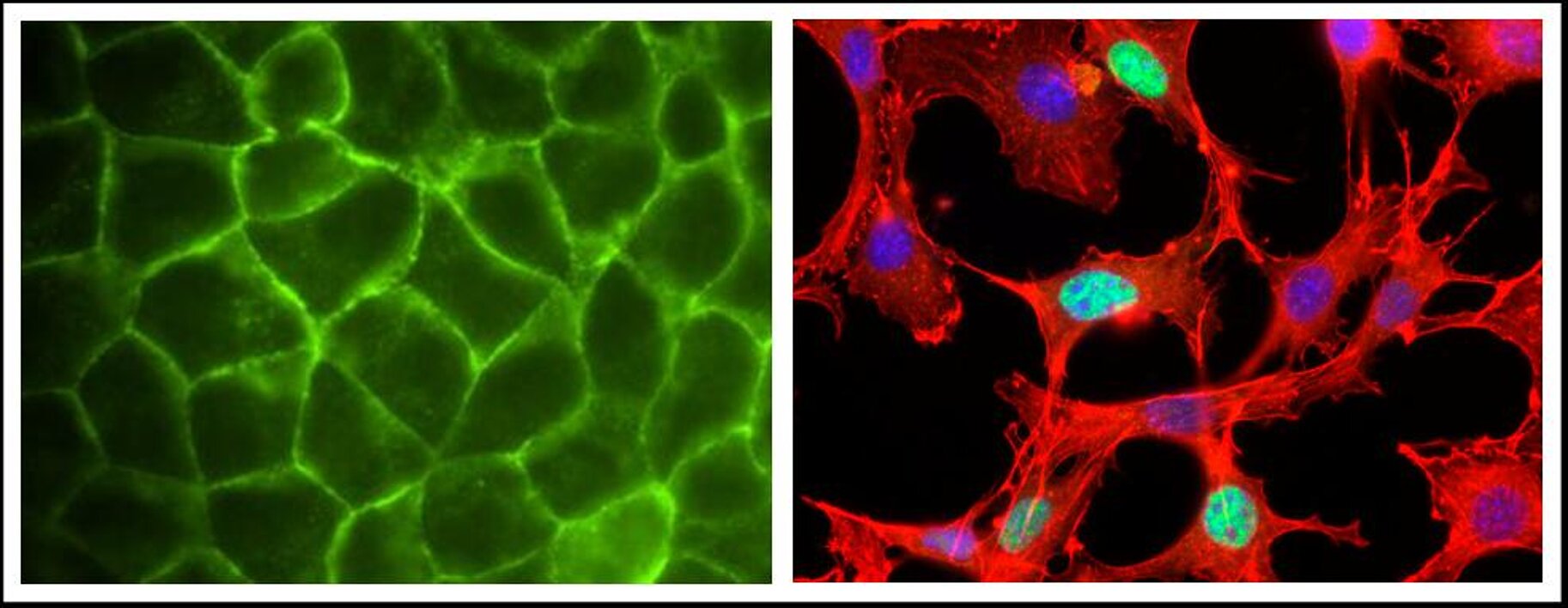
Precise regulation of gene expression in time and space is thought to be critical for cellular function, and many involved transcription factors have been characterised. The cytoskeleton plays an equally important role for the morphology and motility of cells, but the connections between the cytoskeleton and transcriptional control have remained unclear. Recently, we found that a major building block of the cytoskeleton, the monomeric G-actin, is a direct signalling intermediate which connects cytoskeletal dynamics and transcriptional control.
The regulation of gene expression through monomeric G-actin is at the core of our studies. This pathway activates serum response factor through its transcriptional coactivator MAL/MRTF, thereby inducing a distinct subset of target genes. Ras-independent, Rho/Rac dependent target genes require MAL binding to SRF on their promoters for efficient activation. This is inhibited by G-actin, which binds directly to MAL and thereby blocks it. However, the plethora of G-actin regulated genes is unknown, as well as their overall cellular function.
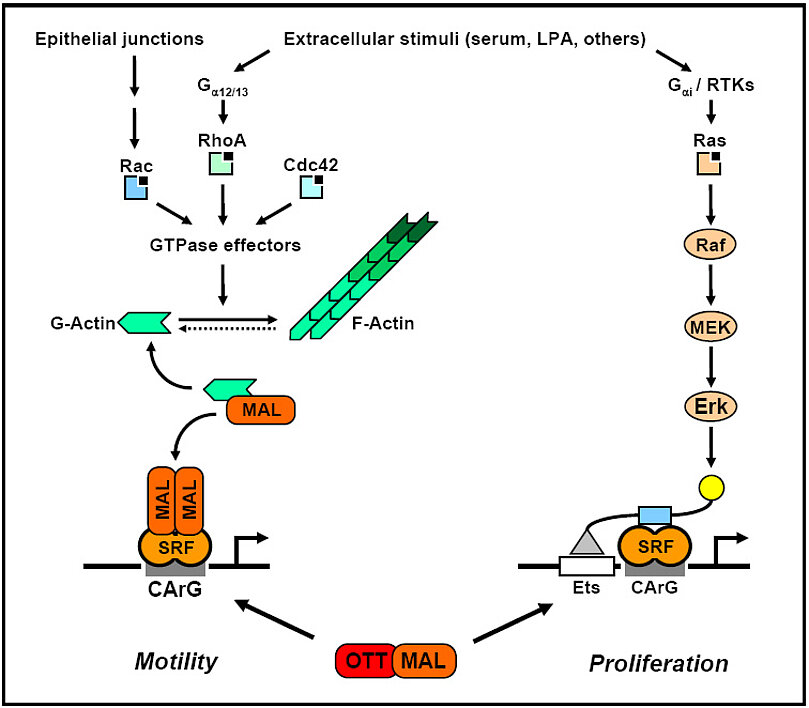
We aim at understanding the regulation and function of the actin-dependent gene expression. Therefore, we are looking for inducers of actin-MAL signalling in largely serum-insensitive epithelial cells, since the regulation of MAL has previously been analysed in mesodermal cells only. Importantly, epithelial tissues are characterised by specialised cell-cell junctions, which disassemble during epithelial-mesenchymal transition (EMT). We could show that junctional disassembly and EMT, which causes actin reorganisation, specifically activates MAL and SRF in epithelia (Busche et al, 2008; 2010). In contrast to serum-stimulated fibroblasts, which require the GTPase RhoA, we found that signalling in epithelia occurs via Rac. The distinct upstream signalling pathways merged at the level of G-actin to activate MAL and SRF, and subsequent target genes. We will continue this project by analysing which junctional components trigger actin-MAL signalling.
MAL was originally identified in a leukemogenic translocation, t(1,22), which results in the expression of the fusion protein OTT-MAL. We analysed this fusion protein with regard to its transcriptional activity and found that OTT-MAL is a deregulated, constitutive activator of SRF-mediated gene expression (Descot et al, 2008). In contrast to the tightly controlled MAL protein, OTT‑MAL is not regulated via Rho-actin signalling and cannot bind monomeric actin. Hence it is always nuclear and activates reporters and endogenous genes. OTT-MAL also causes a delayed induction of the MAL-independent, TCF-dependent target genes c‑fos and egr‑1, and the MAPK/Erk pathway. Our data suggest that the deregulated activation of MAL-dependent and ‑independent promoters results in specific functions of OTT‑MAL.
In order to understand the overall biological function of actin-regulated gene expression and unravel the underlying transcriptional networks, we are frequently using functional genomics. We perform genome-wide expression analysis with affymetrix microarrays using various cell types. As yet, we identified approximately 200 genes directly regulated by G-actin (Descot et al, 2009). Strikingly, they included many previously known MAL-dependent SRF targets, but largely excluded MAPK-regulated genes such as the prototypical growth-promoting oncogenes c-fos and egr-1, which nicely confirmed the suitability of our screening approaches. The unknown targets included many genes with a role in migration, adhesion and motility (Leitner et al, 2012). This suggests the existence of a feed forward loop towards a highly migratory phenotype.
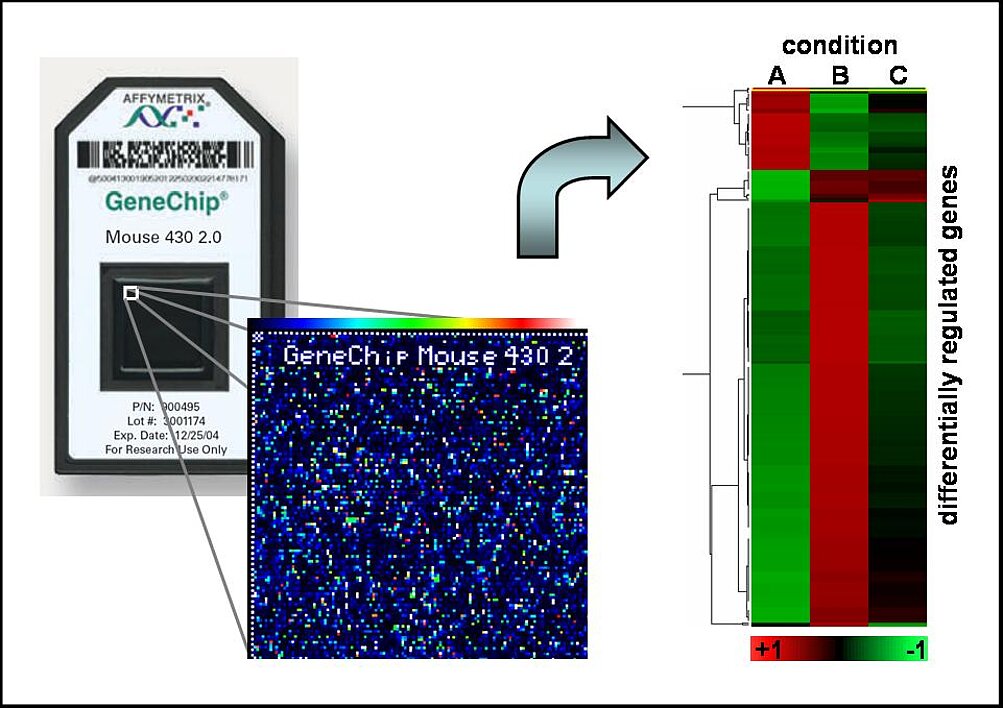
We will continue analysing the ins and outs of actin-regulated gene expression. Therefore we will use suitable tissue culture models for EMT and different modes of migration as well as animal models. In support, novel target genes will be functionally and molecularly characterised, and we will investigate the molecular details of the actin-MAL dissociation. The results will then be applied to a better understanding of G-actin regulated transcriptional networks in physiology and pathology.
We are constantly looking for highly skilled and motivated MSc students, PhD students and Postdocs who are specifically interested in doing research in the above field. Complete applications including Lebenslauf/CV, certificates, research abstracts / list of publications and the names, phone and E-mail addresses of 1-2 academic referees should be send by E-mail.
Research is mainly funded by extramural sources. Current third party-funding includes generous grants from the following:
Scientific cooperation with:
- Core Facilities of the Medical Faculty
- Charles Tanford Protein Centre
- Institute for Biochemistry and Biotechnology
- Universitätsklinik und Poliklinik für Innere Medizin I - Prof. Dr. med. Patrick Michl
- Universitätsklinik und Poliklinik für Innere Medizin IV - apl. Prof. Dr. med. Lutz P. Müller



